- HOME
- VENUE
- RSVP
- REGISTRY
- CONTACT
- Metaclean reviews
- Gta san andreas free downloads for pc
- 2d image 3d live visuals
- Sims 4 city living gameplay
- Managing windows r with vbscript and wmi
- Citrix workspace feature matrix
- Pianoteq 5 promotional code
- Latest radha soami shabad 2015
- Mac two sided printing greyed out hp laserjet 1022
- Bobby fischer my 60 memorable games hardcover
- Download mapinfo 10
- Descargar autocad 2008 64 bits
- Gartner hype cycle robotic process automation
- What frequency mode should wxtoimg for noaa
- How to reset verizon mifi 4510l
- Minecraft 1-12-2 mods intelligent backpacks
- Usb burning tool mxq pro 4k windows 7
- Interspire email marketer api
- Professional dpf egr remover software 3-0 download
- Quik gopro
- Chem draw nmr
- Soundtoys sale 2020
- We are extended music video ice age 4
- Hp designjet 510 driver download mac
- Abby fine reader download
- S s s w w w club
First, it utilized a small set of only 12 peaks (6 high- and 6 low-quality), which likely don’t capture the large variation in peak quality observed in LC–MS metabolomics data ( Coble and Fraga 2014 Myers et al. However, this evaluation was limited in several ways. (2014) proposed and evaluated the effectiveness of six quantitative LC–MS peak quality metrics in filtering out low-quality peaks. Some computational methods have been developed to directly assess peak integration.
#Metaclean reviews manual#
Without a way to automatically and objectively assess integration quality in LC–MS data, manual quality assessment, which is both time-consuming and subjective, is the only way to ensure that poor peak integrations do not propagate to downstream analyses ( Schiffman et al. However, WiPP is only applicable to GC–MS data, and does not consider shifts in retention time that are common to untargeted data generated with LC. 2019) is the only available tool that assesses integration quality and automatically filters any poorly integrated peaks. 2019), but they do not provide a means to evaluate the integration quality following pre-processing. 2015) and enhancing the detection of low abundance metabolites ( Chong et al.
#Metaclean reviews software#
Each of these tools improves the performance of pre-processing software in various ways, such as increasing the number of “reliable” peaks ( Libiseller et al. 2013), and warpgroup utilizes subregion and consensus integration bound detection ( Mahieu et al.

2015), xMSanalyzer improves peak detection by merging multiple datasets produced by varying the parameters of methods from XCMS and apLCMS ( Uppal et al. For example, IPO optimizes XCMS parameters using isotope data ( Libiseller et al. Several tools have been developed to reduce the number of low-quality integrations in LC–MS metabolomics data mainly through optimizations of the pre-processing software mentioned above. This can lead to spurious conclusions from downstream data analyses, if the peaks are not manually curated. The result is that pre-processed output tables, even after traditional filtering approaches, can still contain incorrect abundance values. Particularly for RSD filtering, integration of noise, multiple peaks, or partial peak integrations can be reproducible in sequential measurements of the same sample, but they may not be reliable across many independent biological samples. While this reduces the total number of peaks to a few thousand, and improves the ratio of high-quality peaks to low-quality ones in the final dataset, the latter still usually remain in large numbers ( Schiffman et al.
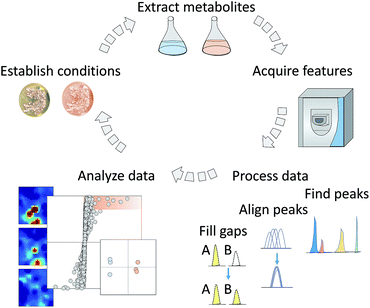
In particular, the most common filtering method, relative standard deviation (RSD) calculated on peaks in a routinely injected pooled quality control (QC) sample, retains peaks that are within a typical threshold, e.g., RSD < 30% ( Broadhurst et al. Subsequent filtering strategies based on pre-determined thresholds on metrics, such as mean/median value across samples, variability across biological samples, and levels of missing values, are routinely employed to remove noisy peaks ( Chong et al. These include large variations in peak detection across software, high prevalence of false positive peaks, and poor integration of identified peaks ( Coble and Fraga 2014 Myers et al. However, despite the availability and diversity of pre-processing software, significant challenges in detecting and integrating peaks persist. A variety of pre-processing software exist for LC–MS data, including both commercial, such as MassHunter Profinder (Agilent), Progenesis QI (Waters), and Compound Discoverer (Thermo), as well as open-source, such as XCMS ( Smith et al. This process yields a table of features, or peaks, (of specific m/z and rt), and their respective abundances in every sample ( Dunn et al. Prior to downstream analyses, these data are generally pre-processed using software to identify peaks representing chemicals in each sample, perform retention time correction, and then group similar peaks across all the samples. A commonly used tool for untargeted metabolomics analyses is liquid chromatography paired with high-resolution mass spectrometry (LC–MS), which generates raw data for chemicals in three dimensions: mass-to-charge ratio ( m/z), retention time (rt) and abundance.